Selection against carbonic anhydrase IX
Once the library has been purified and tested to be suitable for sequencing, selection are carried out under several conditions. Here we show selection of DEL 115 under one condition:

The selection starts by immobilising the protein on magnetic beads and the pre-clearing of the library, consisting in removing bead-binding compounds by incubating naked beads with DEL, which is done in parallel. The resulting pre-cleared library is then incubated with the immobilised protein. After the washes to remove weak binders, the stronger binders are collected by denaturing the protein; this is the 1st round of selection. In this case, the output of the first round is incubated with a new batch of immobilised protein and the second round of selection is collected.
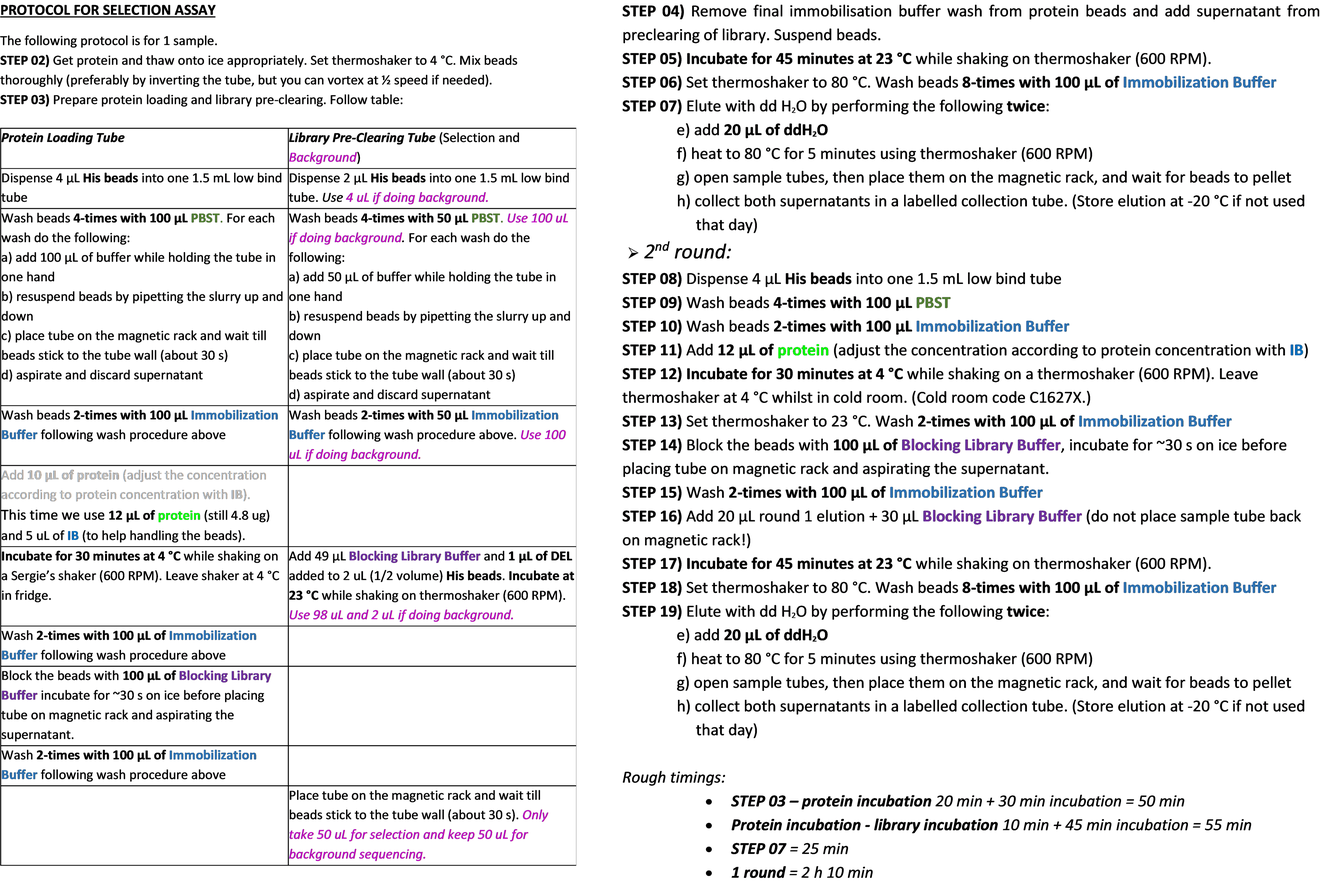
The output of each round of selection is amplified through PCR and sequencing adapters attached. After purification, this PCR amplicons are sent for sequencing.

Sequencing data
The sequencing data is in a FASTQ file format that needs to be deconvoluted and manipulated before data analysis can be carried out to determine number of counts and enrichment factors of all binders. We used KNIME to design workflows that allow for both deconvolution and analysis of the sequencing data.
When looking at enrichment, we look at the increase of a compound or relevant building block (the one with potential interaction with protein, BB3) with respect the background or input library.
In this case study, sequencing of the pre-cleared library provided the background of DEL used as input for the selection. Table X shows the counts and enrichment factors for the top 9 binders in Background 25.
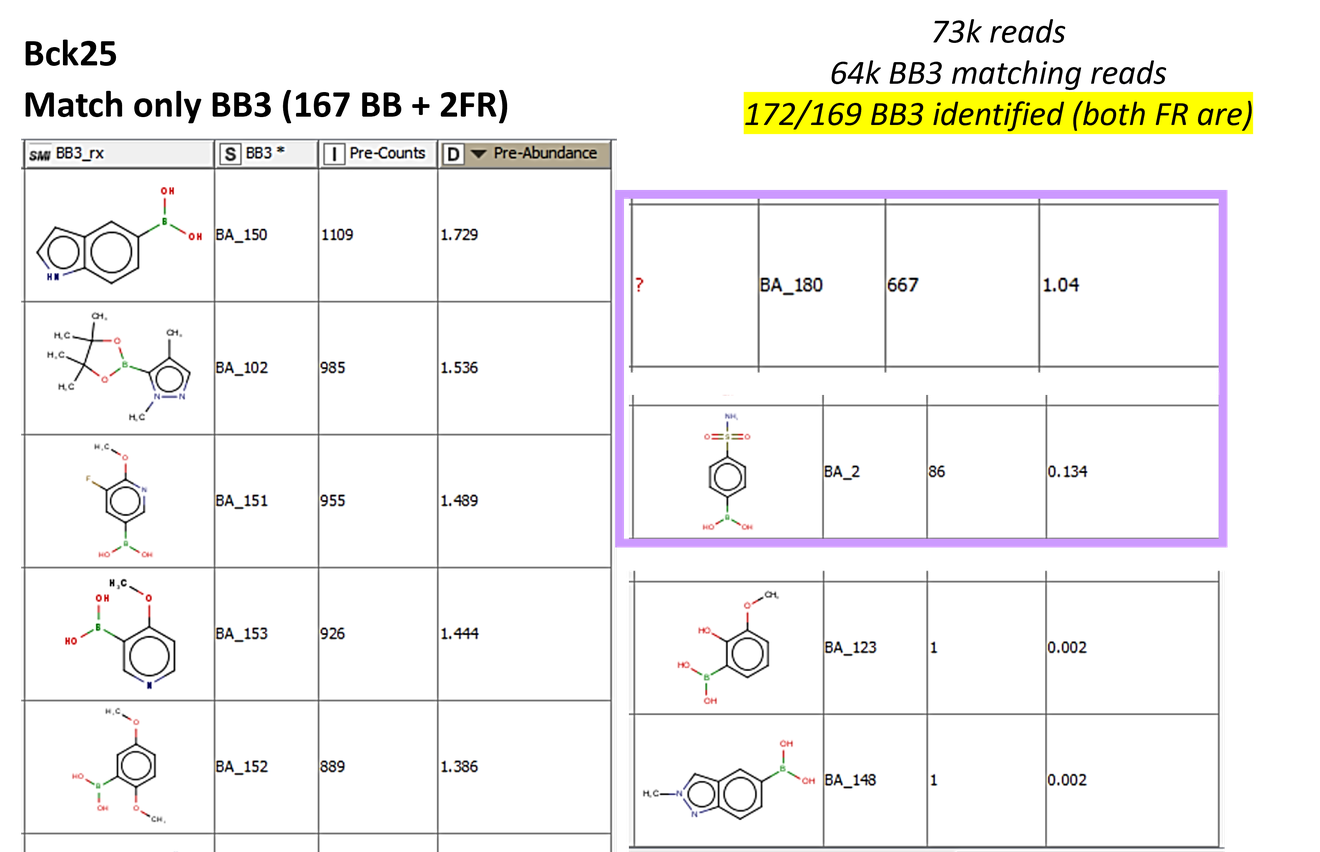
The following histogram shows the number of counts for each BB3 identified in the background 25.
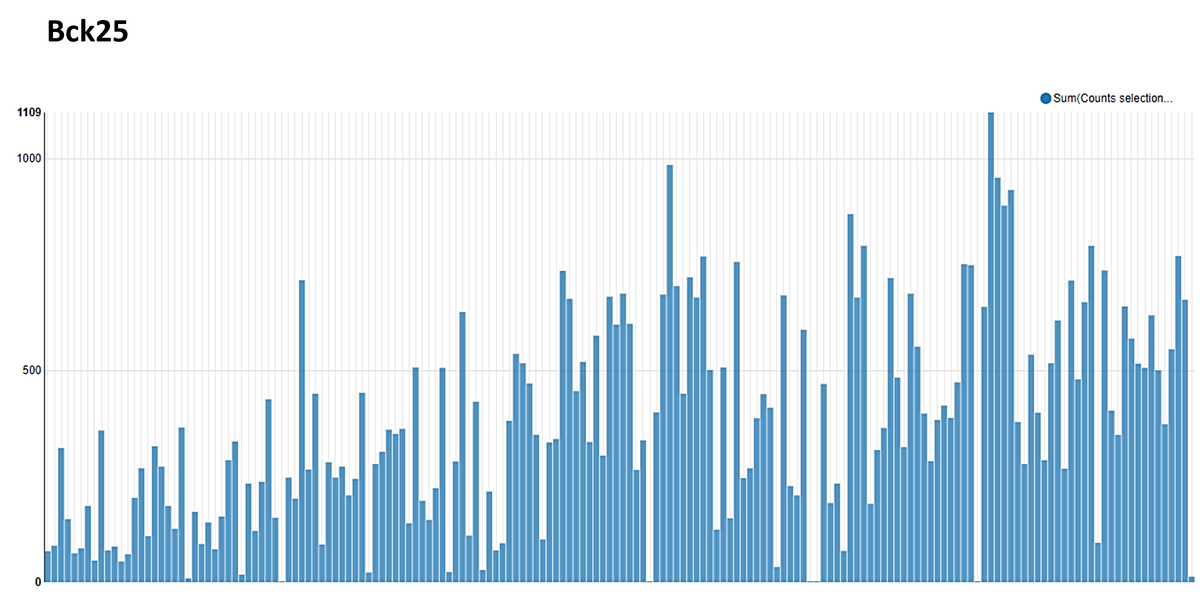
1st round of selection, SELE25-1
Project
- Project ID
- Case Study Test
User
- Name
- Sergie Celis
- Institution
- University of Manchester
Protein
- ID (UniProt code)
- R5555
- Source
- Asp 414
- Supplier
- Acro Biosystems
- Stock (µg)
- 12.0
- State
- lyophilised
- Protein Buffer PB
- 10 mM MES, 200 mM NaCl, pH 6.5
- Reconstitution solvent
- water
- Reconstitution volume (µL)
- 0.3

